木聚糖具有多个不同的取代基,是一种结构极其复杂、分化度很高的多聚五碳糖,主要存在于植物细胞的细胞壁中,是植物细胞半纤维素的重要组成成分[1]。木聚糖的生物降解需要多种酶的协同作用,这些能降解木聚糖的水解酶统称为木聚糖酶。木聚糖酶有一个复杂的酶系统[2],其中最关键的一类木聚糖酶为β-1,4-内切-D-木聚糖酶(endoβ-1,4-D-xylanase)[EC3.2.1.8],该木聚糖酶能降解木聚糖的主链结构[3-4]。
木聚糖酶来源广泛,包括陆地植物组织、细菌、真菌和原生动物等,但大部分来源于陆地。海洋中物种丰富,种类繁多,是人们赖以生存的物质基础[5-6]。海洋微生物产生的木聚糖酶与陆地微生物产生的酶相比具有其特殊性,这些特性使得海洋酶在基础研究和工业应用等方面具有巨大价值,近些年对海洋微生物木聚糖酶的研究成为现在微生物木聚糖酶研究的热点。
1 产木聚糖酶海洋微生物
目前,海洋木聚糖酶主要从细菌[7-8]、真菌[9]以及放线菌[10-11]中分离得到,且细菌产木聚糖酶通常是首选。
海洋来源的木聚糖酶具有嗜盐、嗜热、厌氧等特殊性。从海洋细菌中获得的木聚糖酶大多数为β-1,4-内切-D-木聚糖酶,这些木聚糖酶大多数属于糖苷水解酶家族10和11,但也有属于糖苷水解酶(glucoside hydrolase,GH)家族8、26和30,分子质量大多数在22~50 kDa。最近报道的产木聚糖酶的海洋细菌、木聚糖酶基因、GH家族及分子质量见表1。
表1 产木聚糖酶的海洋细菌
Table 1 Xylanase-producing marine bacteria
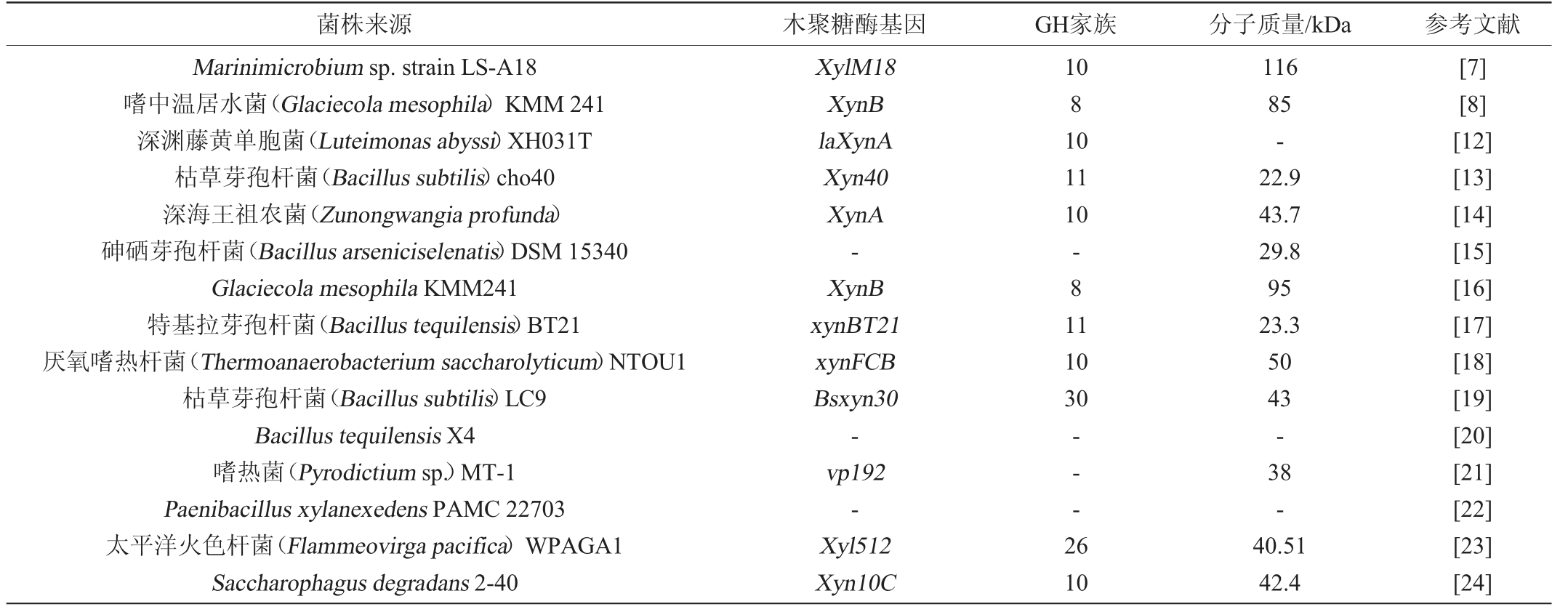
注:“-”表示目前尚未研究清楚。下同。
木聚糖酶可由多种微生物产生,海洋真菌是木聚糖酶的重要来源,但目前对海洋真菌来源的木聚糖酶的研究相对于海洋细菌较少,对其来源的木聚糖酶基因研究尚不清楚。目前,经过筛选已经获得了几种能产生木聚糖酶的海洋真菌见表2。
表2 产木聚糖酶的海洋真菌
Table 2 Xylanase-producing marine fungi
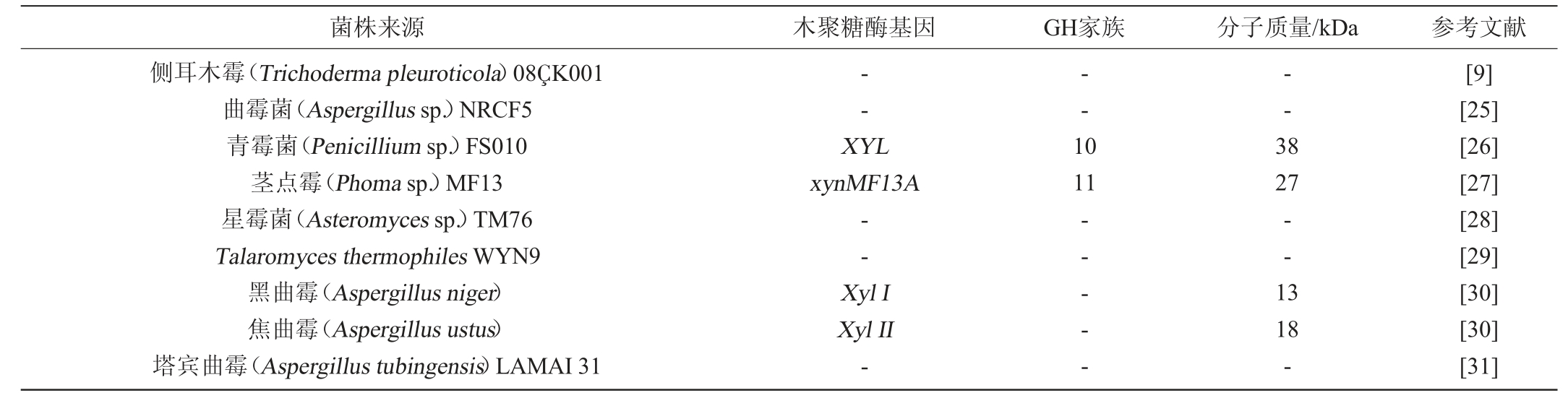
许多海洋放线菌除了产生负责天然产物生物合成的独特酶外,还产生能够降解各种生物聚合物(如木聚糖、纤维素和甲壳素)的水解酶。产木聚糖酶的海洋放线菌见表3。
表3 产木聚糖酶的海洋放线菌
Table 3 Xylanase-producing marine actinomycetes
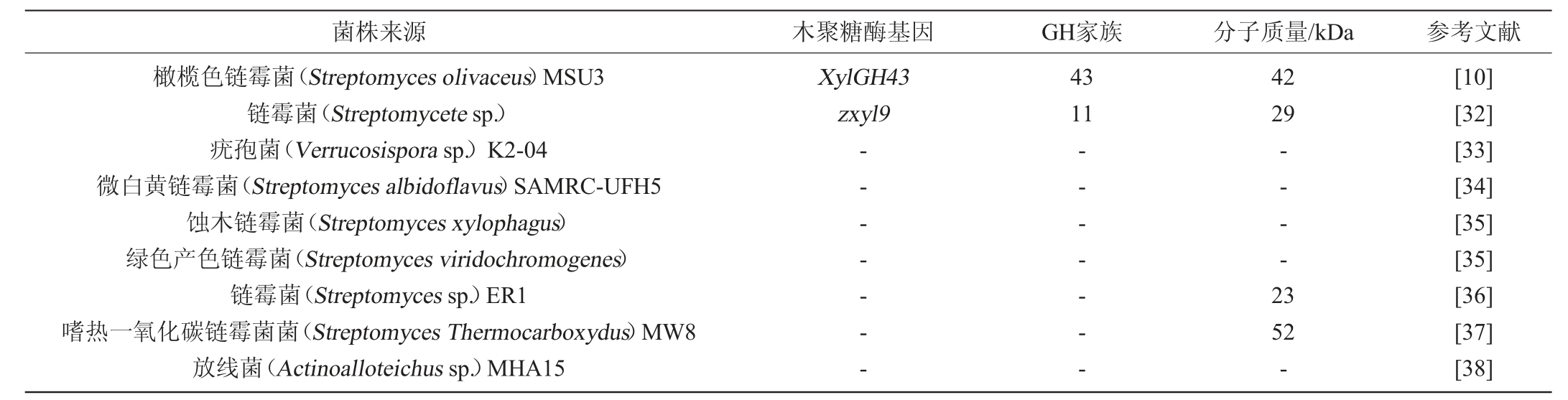

2 产木聚糖酶菌株的选育
海洋微生物的基因组资源特异性丰富,通过对海洋微生物来源的菌株选育,能够筛选出具有特异性的产木聚糖酶的海洋菌株,对这些菌株的基因序列和功能进行分析和鉴定,能够得到嗜盐、嗜热、厌氧等不同功能的木聚糖酶基因。如黄小云等[39]在多种海洋来源的样品中筛选出了38株具有产木聚糖酶能力的菌株,其中菌株B659产酶能力最高,酶活力为525.3 U/mL;YOPI Y等[20]从20株海洋芽孢杆菌分离株中分离、鉴定得到产甘露聚糖酶、木聚糖酶和纤维素酶的海洋芽孢杆菌菌株,分别为枯草芽孢杆菌(Bacillus subtilis)、特基拉芽孢杆菌(Bacillus tequilensis)和蜡样芽孢杆菌(Bacillus cereus);HOU Y H等[26]从黄海沉积物中分离到1株冷适应青霉菌(Penicillium)FS010,并从该菌中获得冷活性木聚糖酶XYL。
3 木聚糖酶基因的克隆及分子改造
产木聚糖酶菌株筛选的环境不同,获得的菌株生长条件及产酶活力也存在很大差异。对菌株进行基因克隆与改造,能优化菌株的培养条件,提高其产酶能力。GUO B等[16]从嗜中稳居水菌(Glaciecola mesophila)KMM241中克隆到一个新的木聚糖酶基因xynB,并成功在大肠杆菌(Escherichia coli)中获得异源表达;GUO Y等[19]将内切木聚糖酶基因克隆到枯草芽孢杆菌WB800中进行蛋白表达和鉴定,获得重组木聚糖酶活性为55 U/mL,比原始菌株酶活性提高了14.5倍;AHMED M A等[25]以Aspergillus sp.NRCF5为出发菌株,利用遗传变异和原生质体融合的方法,通过4轮基因重组获得了高产木聚糖酶的重组菌;刘卓[35]通过锌转录因子与木聚糖酶相关基因启动子上游区域结合,调节了木聚糖酶的产生,证明了人工锌指蛋白技术和核糖体工程技术在改良放线菌菌种方面是可行的,为探索放线菌木聚糖酶的合成规律,代谢工程转化海洋酶蛋白奠定了基础。
4 产木聚糖酶菌株发酵工艺的研究
发酵条件也可以影响酶的活力。YU H等[7]从Marinimicrobium sp.LS-A18中克隆得到木聚糖酶(XylM18)基因,在大肠杆菌BL21(DE3)中异源表达,工程菌发酵工艺优化,重组XylM18对木聚糖的表观米氏常数(Km)和Vmax分别为1.60 mg/mL和130.1 U/mg;刘斌等[28]研究结果显示,Asteromyces sp.TM76发酵工艺优化后,木聚糖酶的最大酶活力为470.5 U/mL;DOS SANTOS J A等[31]通过对493株海洋微生物筛选及相应的发酵工艺优化,最终确定Aspergillus cf. tubingensis LAMAI 31可作为木聚糖生产的新菌种;EVELYN N等[34]从链霉菌属中获得木聚糖酶生产菌株,产木聚糖酶的最优工艺为pH8.0、培养温度35℃、转速150r/min,木聚糖酶的比酶活力为7 417 U/g;林小洪等[40]从福建平潭海域筛选到1株产木聚糖酶活力较高的菌株B659,经16S rDNA基因序列分析鉴定为同温层芽胞杆菌(Stratosphericbacillus),通过发酵工艺的优化,木聚糖酶的活性达到了2 838 U/mL,为优化前的3.4倍,发酵周期也从72 h缩短至30 h。
5 木聚糖酶在工业中的应用
木聚糖酶被广泛应用于饲料加工、制浆造纸、纺织以及食品等众多工业领域,尤其是在造纸以及纸浆漂白工业中,木聚糖酶的应用更加广泛[41]。RAGHUKUMAR C等[30]首次报道了一种耐高温、无纤维素酶的碱性木聚糖酶的粗培滤液在纸浆生物浸出中的潜在应用;KHANDEPARKER R等[17]研究发现,木聚糖酶XynBT21具有从麦麸中生产木二糖的潜力;WU J等[27]认为木聚糖酶rXynMF13A对馒头的生产有一定的促进作用,在烘焙、海产品加工中具有应用潜力;ROSMINE E等[36]研究发现,木聚糖酶对橙汁、菠萝汁的澄清率分别为20.87%、27.89%,因此链霉菌ER1产生的木聚糖酶可用于果汁澄清;CHENG A C等[42]最近研究证实,在饲料中添加外源木聚糖酶可以改善罗非鱼对养分的利用,从而促进鱼类的生长;DJOHAN A C等[43]使用本土海洋细菌沙福芽孢杆菌(Bacillus safensis)LBF002的木聚糖酶可将稻草废料转化为还原糖低聚木糖,如木二糖、木己糖和木糖,将该木聚糖酶与含有25 g/L稻草的反应混合物在50 ℃孵育4 h,可获得1 g/L的还原糖。
6 结论与展望
海洋木聚糖酶的来源广泛,在细菌、真菌以及放线菌中都有木聚糖酶被筛选出来。木聚糖酶的应用主要体现在:在饲料工业中,水解饲料里的木聚糖成分,提高饲料的营养价值和利用率;在造纸工业中,用于纸浆的漂白,减少环境的污染;在食品行业上,用来生产功能性木寡糖,改善面包品质等。海洋木聚糖酶的产生环境非常广泛,包括酸性到碱性环境以及中度嗜冷到中度嗜热环境等[44]。此外,大多数海洋木聚糖酶都是嗜盐的,有些可以承受金属离子的存在,这些金属离子往往对包括陆源木聚糖酶在内的其他酶有抑制作用[45],这些特征突出了海洋木聚糖酶在工业应用领域的特殊潜力。对海洋微生物的研究,可能会分离出更多催化潜力更强的海洋木聚糖酶。
[1]王晶,董翔,覃瑞,等.新疆胡杨树皮提取木聚糖的研究[J].湖北农业科学,2018,57(3):75-77.
[2]TAN Z B,TANG C D,WU M C,et al.Exploration of disulfide bridge and N-glycosylation contributing to high thermostability of a hybrid xylanase[J].Protein Pept Lett,2014,21(7):657-662.
[3]LI G Q,CHEN X J,ZHOU X,et al.Improvement of GH10 family xylanase thermostability by introducing of an extra α-helix at the C-terminal[J].Biochem Biophys Res Commun,2019,515(3):417-422.
[4] DUTTA B,BANERJEE A,CHAKRABORTY P.Studies on bacterial xylanase enzyme:structural and functional insight[J].J Genet Eng Biotechnol,2018,16(2):749-756.
[5] ROMANO G,COSTANTINI M,SANSONE C,et al.Marine microorganisms as a promising and sustainable source of bioactive molecules[J].Mar Environ Res,2017,128:58-69.
[6]GLÖCKNER F O,JOINT I.Marine microbial genomics in Europe:current status and perspectives[J].Microb Biotechnol,2010,3(5):523-530.
[7]YU H,ZHAO S X,FAN Y Q,et al.Cloning and heterologous expression of a novel halo/alkali-stable multi-domain xylanase (XylM18) from a marine bacterium Marinimicrobium sp.strain LS-A18[J].Appl Microbiol Biotechnol,2019,103(21-22):8899-8909.
[8]CHEN X L,ZHAO F,YUE Y S,et al.A new group of modular xylanases in glycoside hydrolase family 8 from marine bacteria[J].Appl Environ Microbiol,2018,84(23):e01785-e01718.
[9]KORKMAZ M N,OZDEMIR S C,UZEL A.Xylanase production from marine derived Trichoderma pleuroticola 08ÇK001 strain isolated from Mediterranean coastal sediments[J].J Basic Microbiol,2017,57(10):839-851.
[10] SANJIVKUMAR M,SILAMBARASAN T,PALAVESAM A,et al.Biosynthesis,purification and characterization of β-1,4-xylanase from a novel mangrove associated actinobacterium Streptomyces olivaceus(MSU3) and its applications[J].Protein Expr Purif,2017,130:1-12.
[11] MUDDALA R,ACOSTA J A M,BARBOSA L C A,et al.Synthesis of the marine myxobacterial antibiotic enhygrolide A[J].J Nat Prod,2017,80(7):2166-2169.
[12] HAN Z,SHANG G F,YANG J.Characterization of a novel cold-active xylanasefrom Luteimonasspecies[J].World J Microbiol Biotechnol,2018,34(8):123.
[13] KHANDEPARKER R,VERMA P,DEOBAGKAR D.A novel halotolerant xylanase from marine isolate Bacillus subtilis cho40:gene cloning and sequencing[J].N Biotechnol,2011,28(6):814-821.
[14]LIU X,HUANG Z,HUANG X,et al.Cloning,expression and characterization of a novel cold-active and halophilic xylanase from Zunongwangia profunda[J].Extremophiles,2014,18(2):441-450.
[15]KAMBLE R D,JADHAV A R.Isolation,purification,and characterization of xylanase produced by a new species of Bacillus in solid state fermentation[J].Int J Microbiol,2012:683193.
[16]GUO B,LI P Y,YUE Y S,et al.Gene cloning,expression and characterization of a novel xylanase from the marine bacterium,Glaciecola mesophila KMM241[J].Mar Drugs,2013,11(4):1173-1187.
[17] KHANDEPARKER R,PARAB P,AMBERKAR U.Recombinant xylanase from Bacillus tequilensis BT21:Biochemical characterisation and its application in the production of xylobiose from agricultural residues[J].Food Technol Biotechnol,2017,55(2):164-172.
[18] HUNG K S,HUNG S M,TOU W S,et al.Characterization of a novel GH10 thermostable,halophilic xylanase from the marine bacterium Thermoanaerobacterium saccharolyticum NTOU1[J].Process Biochem,2011(46):1257-1263.
[19]GUO Y,GAO Z,XU J,et al.A family 30 glucurono-xylanase from Bacillus subtilis LC9:expression,characterization and its application in Chinese bread making[J].Int J Biol Macromol,2018,117:377-384.
[20]YOPI Y,DJOHAN A C,RAHMANI N,et al.Isolation and characterization of mannanase,xylanase,and cellulase from marine bacteria Bacillus sp.[J].J Natural Products Biochem,2017,15(1):15-20.
[21]吴穗洁.深海嗜热细菌热稳定木聚糖酶的性质及噬菌体GSVE1 功能基因的研究[D].厦门:厦门大学,2007.
[22] HWANG J,SHIN S C,HAN J W,et al.Complete genome sequence of Paenibacillus xylanexedens PAMC 22703,a xylan-degrading bacterium[J].Mar Drugs,2020,55:100788.
[23] CAI Z W,GE H H,YI Z W,et al.Characterization of a novel psychrophilic and halophilic β-1,3-xylanase from deep-sea bacterium, Flammeovirga pacifica strain WPAGA1[J].Int J Biol Macromol,2018,118:2176-2184.
[24]KO J K,KO H,KIM K H,et al.Characterization of the biochemical properties of recombinant Xyn10C from a marine bacterium, Saccharophagus degradans 2-40[J].Bioproc Biosyst Eng,2016,39(4):677-684.
[25] AHMED M A,BONDKLY E L.Molecular identification using ITS sequences and genome shuffling to improve 2-deoxyglucose tolerance and xylanase activity of marine derived fungus,Aspergillus sp.NRCF5[J].Appl Biochem Biotechnol,2012,167(8):2160-2173.
[26]HOU Y H,WANG T H,LONG H,et al.Novel cold-adaptive Penicillium strain FS010 secreting thermolabile xylanase isolated from Yellow Sea[J].Acta Biochim Biophys Sin,2006,38(2):142-149.
[27]WU J,QIU C,REN Y,et al.Novel salttolerant xylanase from a mangrove isolated fungus Phoma sp.MF13 and its application in Chinese steamed bread[J].ACS Omega,2018,3(4):3708-3716.
[28]刘斌,李正魁,蔡敬民,等.海洋真菌Asteromyces sp.TM76 木聚糖酶生产初步研究[J].工业微生物杂志,2005,35(3):6-9.
[29]吴亚宁,曾艾伦,张玉媚,等.一株产木聚糖酶嗜热真菌的分子鉴定及酶学性质研究[J].食品科技,2013,38(9):20-24.
[30] RAGHUKUMAR C,MURALEEDHARAN U,GAUD V R,et al.Xylanases of marine fungi of potential use for biobleaching of paper pulp[J].J Ind Microbiol Biotechnol,2004,31(9):433-441.
[31] DOS SANTOS J A,VIEIRA J M F,VIDEIRA A,et al.Marine-derived fungus Aspergillus cf.tubingensis LAMAI 31:A new genetic resource for xylanase production[J].AMB Express,2016,6(1):25.
[32]王吟,杨艳燕,肖静,等.链霉菌zxy19 木聚糖酶酶学性质及酶基因克隆[J].微生物学通报,2008,35(11):1681-1685.
[33]OMAR S M,FAROUK N M,MALEK N A,et al.Verrucosispora sp.K2-04,potential xylanase producer from Kuantan Mangrove forest sediment[J].Int J Food Eng,2017,3(2):165-168.
[34]FATOKUN E N,NWODO U U,OKOH A I.Classical optimization of cellulase and xylanase production by a marine Streptomyces species[J].Appl Sci,2016,6(10):286.
[35]刘卓.海洋来源产木聚糖酶的链霉菌菌种筛选与代谢工程改造[D].大连:大连理工大学,2013.
[36]ROSMINE E,SAINJAN N C,SILVESTER R,et al.Statistical optimisation of xylanase production by estuarine Streptomyces sp.and its application in clarification of fruit juice[J].J Genet Eng Biotechnol,2017,15(2):393-401.
[37]CHI W J,LIM J H,PARK D P,et al.Production and characterization of a thermostable endo-type β-xylanase produced by a newly-isolated Streptomyces thermocarboxydus subspecies MW8 strain from Jeju Island[J].Process Biochem,2013,48(11):1736-1743.
[38]RAJAGOPAL G,KANNAN S.Systematic characterization of potential cellulolytic marine actinobacteria Actinoalloteichus sp.MHA15[J].Biotechnol Rep,2016,13:30-36.
[39]黄小云,林娟,林小洪,等.产木聚糖酶海洋微生物的筛选与诱变育种[J].福州大学学报(自然科学版),2015,43(5):715-721.
[40]林小洪,林娟,王国增,等.海洋来源芽孢杆菌产木聚糖酶的条件研究[J].中国食品学报,2015,15(11):83-90.
[41]QESHMI F I,HOMAEI A,FERNANDES P,et al.Xylanases from marine microorganisms:A brief overview on scope,sources,features and potential applications[J].BBA-Proteins Proteom,2020,1868(2):140312.
[42]CHENG A C,CHI C C,LIU C H.Effects of X-xylanase incorporated in the diet on the growth performance of red tilapia(Oreochromis sp.)[J].J Fish Sci,2018(12):33-38.
[43]DJOHAN A C,PERWITASARI U,YOPI Y.Saccharification waste biomass rice straw IR-64 by using xylanase from indigenous marine bacteria Bacillus safensis LBF-002[J].Int J Adv Sci,2016(6):40-44.
[44] TIGHE S,AFSHINNEKOO E,ROCK TM,et al.Genomic methods and microbiological technologies for profiling novel and extreme environments for the extreme microbiome project(XMP)[J].J Biomol Tech,2017,28(1):31-39.
[45]KAMENNAYA N A,GERAKI K,SCANLAN D J,et al.Accumulation of ambient phosphate into the periplasm of marine bacteria is proton motive force dependent[J].Nat Commun,2020,11(1):1-13.